Sequencing Methods
Sanger sequencing
From Wikipedia, the free encyclopedia
Sanger sequencing is a method of DNA sequencing first commercialized by Applied Biosystems, based on the selective incorporation of chain-terminating dideoxynucleotides by DNA polymerase during in vitro DNA replication.[1][2] Developed by Frederick Sanger and colleagues in 1977, it was the most widely used sequencing method for approximately 40 years. More recently, higher volume Sanger sequencing has been supplanted by "Next-Gen" sequencing methods, especially for large-scale, automated genome analyses. However, the Sanger method remains in wide use, for smaller-scale projects, validation of Next-Gen results and for obtaining especially long contiguous DNA sequence reads (> 500 nucleotides).
Method
The classical chain-termination method requires a single-stranded DNA template, a DNA primer, a DNA polymerase, normal deoxynucleosidetriphosphates (dNTPs), and modified di-deoxynucleotidetriphosphates (ddNTPs), the latter of which terminate DNA strand elongation. These chain-terminating nucleotides lack a 3'-OH group required for the formation of a phosphodiester bond between two nucleotides, causing DNA polymerase to cease extension of DNA when a modified ddNTP is incorporated. The ddNTPs may be radioactively or fluorescently labelled for detection in automated sequencing machines.
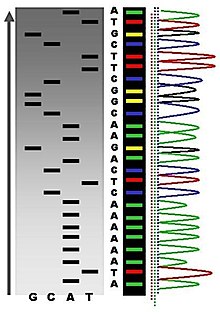
The DNA sample is divided into four separate sequencing reactions, containing all four of the standard deoxynucleotides (dATP, dGTP, dCTP and dTTP) and the DNA polymerase. To each reaction is added only one of the four dideoxynucleotides (ddATP, ddGTP, ddCTP, or ddTTP), while the other added nucleotides are ordinary ones. The dideoxynucleotide concentration should be approximately 100-fold lower than that of the corresponding deoxynucleotide (e.g. 0.005mM ddTTP : 0.5mM dTTP) to allow enough fragments to be produced while still transcribing the complete sequence.[2] Putting it in a more sensible order, four separate reactions are needed in this process to test all four ddNTPs. Following rounds of template DNA extension from the bound primer, the resulting DNA fragments are heat denatured and separated by size using gel electrophoresis. In the original publication of 1977,[2] the formation of base-paired loops of ssDNA was a cause of serious difficulty in resolving bands at some locations. This is frequently performed using a denaturing polyacrylamide-urea gel with each of the four reactions run in one of four individual lanes (lanes A, T, G, C). The DNA bands may then be visualized by autoradiography or UV light and the DNA sequence can be directly read off the X-ray film or gel image.
Part of a radioactively labelled sequencing gel
In the image on the right, X-ray film was exposed to the gel, and the dark bands correspond to DNA fragments of different lengths. A dark band in a lane indicates a DNA fragment that is the result of chain termination after incorporation of a dideoxynucleotide (ddATP, ddGTP, ddCTP, or ddTTP). The relative positions of the different bands among the four lanes, from bottom to top, are then used to read the DNA sequence.
DNA fragments are labelled with a radioactive or fluorescent tag on the primer (1), in the new DNA strand with a labeled dNTP, or with a labeled ddNTP.
Technical variations of chain-termination sequencing include tagging with nucleotides containing radioactive phosphorus for radiolabelling, or using a primer labeled at the 5' end with a fluorescent dye. Dye-primer sequencing facilitates reading in an optical system for faster and more economical analysis and automation. The later development by Leroy Hood and coworkers[3][4] of fluorescently labeled ddNTPs and primers set the stage for automated, high-throughput DNA sequencing.
Sequence ladder by radioactive sequencing compared to fluorescent peaks
Chain-termination methods have greatly simplified DNA sequencing. For example, chain-termination-based kits are commercially available that contain the reagents needed for sequencing, pre-aliquoted and ready to use. Limitations include non-specific binding of the primer to the DNA, affecting accurate read-out of the DNA sequence, and DNA secondary structures affecting the fidelity of the sequence.
Dye-terminator sequencing
Capillary electrophoresis
Dye-terminator sequencing utilizes labelling of the chain terminator ddNTPs, which permits sequencing in a single reaction, rather than four reactions as in the labelled-primer method. In dye-terminator sequencing, each of the four dideoxynucleotide chain terminators is labelled with fluorescent dyes, each of which emit light at different wavelengths.
Owing to its greater expediency and speed, dye-terminator sequencing is now the mainstay in automated sequencing. Its limitations include dye effects due to differences in the incorporation of the dye-labelled chain terminators into the DNA fragment, resulting in unequal peak heights and shapes in the electronic DNA sequence trace chromatogram after capillary electrophoresis (see figure to the left).
This problem has been addressed with the use of modified DNA polymerase enzyme systems and dyes that minimize incorporation variability, as well as methods for eliminating "dye blobs". The dye-terminator sequencing method, along with automated high-throughput DNA sequence analyzers, was used for the vast majority of sequencing projects until the introduction of Next Generation Sequencing.
Automation and sample preparation
View of the start of an example dye-terminator read
Automated DNA-sequencing instruments (DNA sequencers) can sequence up to 384 DNA samples in a single batch. Batch runs may occur up to 24 times a day. DNA sequencers separate strands by size (or length) using capillary electrophoresis, they detect and record dye fluorescence, and output data as fluorescent peak trace chromatograms. Sequencing reactions (thermocycling and labelling), cleanup and re-suspension of samples in a buffer solution are performed separately, before loading samples onto the sequencer. A number of commercial and non-commercial software packages can trim low-quality DNA traces automatically. These programs score the quality of each peak and remove low-quality base peaks (which are generally located at the ends of the sequence). The accuracy of such algorithms is inferior to visual examination by a human operator, but is adequate for automated processing of large sequence data sets.
Challenges
Common challenges of DNA sequencing with the Sanger method include poor quality in the first 15-40 bases of the sequence due to primer binding[citation needed] and deteriorating quality of sequencing traces after 700-900 bases. Base calling software such as Phred typically provides an estimate of quality to aid in trimming of low-quality regions of sequences.[5][6]
In cases where DNA fragments are cloned before sequencing, the resulting sequence may contain parts of the cloning vector. In contrast, PCR-based cloning and next-generation sequencing technologies based on pyrosequencing often avoid using cloning vectors. Recently, one-step Sanger sequencing (combined amplification and sequencing) methods such as Ampliseq and SeqSharp have been developed that allow rapid sequencing of target genes without cloning or prior amplification.[7][8]
Current methods can directly sequence only relatively short (300-1000 nucleotides long) DNA fragments in a single reaction. The main obstacle to sequencing DNA fragments above this size limit is insufficient power of separation for resolving large DNA fragments that differ in length by only one nucleotide.
Shotgun sequencing
From Wikipedia, the free encyclopedia
In genetics, shotgun sequencing is a method used for sequencing long DNA strands. It is named by analogy with the rapidly expanding, quasi-random firing pattern of a shotgun.
The chain termination method of DNA sequencing ("Sanger sequencing") can only be used for short DNA strands of 100 to 1000 base pairs. Due to this size limit, longer sequences are subdivided into smaller fragments that can be sequenced separately, and these sequences are assembled to give the overall sequence.
There are two principal methods for this fragmentation and sequencing process. Primer walking (or "chromosome walking") progresses through the entire strand piece by piece, whereas shotgun sequencing is a faster but more complex process that uses random fragments.
In shotgun sequencing,[1][2] DNA is broken up randomly into numerous small segments, which are sequenced using the chain termination method to obtain reads. Multiple overlapping reads for the target DNA are obtained by performing several rounds of this fragmentation and sequencing. Computer programs then use the overlapping ends of different reads to assemble them into a continuous sequence.[1]
Shotgun sequencing was one of the precursor technologies that was responsible for enabling full genome sequencing.
Hierarchical shotgun sequencing
In whole genome shotgun sequencing (top), the entire genome is sheared randomly into small fragments (appropriately sized for sequencing) and then reassembled. In hierarchical shotgun sequencing (bottom), the genome is first broken into larger segments. After the order of these segments is deduced, they are further sheared into fragments appropriately sized for sequencing.
Although shotgun sequencing can in theory be applied to a genome of any size, its direct application to the sequencing of large genomes (for instance, the human genome) was limited until the late 1990s, when technological advances made practical the handling of the vast quantities of complex data involved in the process.[13] Historically, full-genome shotgun sequencing was believed to be limited by both the sheer size of large genomes and by the complexity added by the high percentage of repetitive DNA (greater than 50% for the human genome) present in large genomes.[14] It was not widely accepted that a full-genome shotgun sequence of a large genome would provide reliable data. For these reasons, other strategies that lowered the computational load of sequence assembly had to be utilized before shotgun sequencing was performed.[14] In hierarchical sequencing, also known as top-down sequencing, a low-resolution physical map of the genome is made prior to actual sequencing. From this map, a minimal number of fragments that cover the entire chromosome are selected for sequencing.[15] In this way, the minimum amount of high-throughput sequencing and assembly is required.
The amplified genome is first sheared into larger pieces (50-200kb) and cloned into a bacterial host using BACs or PACs. Because multiple genome copies have been sheared at random, the fragments contained in these clones have different ends, and with enough coverage (see section above) finding a scaffold of BAC contigs that covers the entire genome is theoretically possible. This scaffold is called a tiling path.
A BAC contig that covers the entire genomic area of interest makes up the tiling path.
Once a tiling path has been found, the BACs that form this path are sheared at random into smaller fragments and can be sequenced using the shotgun method on a smaller scale.
Although the full sequences of the BAC contigs is not known, their orientations relative to one another are known. There are several methods for deducing this order and selecting the BACs that make up a tiling path. The general strategy involves identifying the positions of the clones relative to one another and then selecting the least number of clones required to form a contiguous scaffold that covers the entire area of interest. The order of the clones is deduced by determining the way in which they overlap.[16] Overlapping clones can be identified in several ways. A small radioactively or chemically labeled probe containing a sequence-tagged site (STS) can be hybridized onto a microarray upon which the clones are printed.[16] In this way, all the clones that contain a particular sequence in the genome are identified. The end of one of these clones can then be sequenced to yield a new probe and the process repeated in a method called chromosome walking.
Alternatively, the BAC library can be restriction-digested. Two clones that have several fragment sizes in common are inferred to overlap because they contain multiple similarly spaced restriction sites in common.[16] This method of genomic mapping is called restriction fingerprinting because it identifies a set of restriction sites contained in each clone. Once the overlap between the clones has been found and their order relative to the genome known, a scaffold of a minimal subset of these contigs that covers the entire genome is shotgun-sequenced.[15]
Because it involves first creating a low-resolution map of the genome, hierarchical shotgun sequencing is slower than whole-genome shotgun sequencing, but relies less heavily on computer algorithms than whole-genome shotgun sequencing. The process of extensive BAC library creation and tiling path selection, however, make hierarchical shotgun sequencing slow and labor-intensive. Now that the technology is available and the reliability of the data demonstrated,[14] and the speed and cost efficiency of whole-genome shotgun sequencing has made it the primary method for genome sequencing.
Illumina dye sequencing
From Wikipedia, the free encyclopedia
The DNA attaches to the flow cell via complementary sequences. The strand bends over and attaches to a second oligo forming a bridge. A polymerase synthesizes the reverse strand. The two strands release and straighten. Each forms a new bridge (bridge amplification). The result is a cluster of DNA forward and reverse strands clones.
Illumina dye sequencing is a technique used to determine the series of base pairs in DNA, also known as DNA sequencing. The reversible terminated chemistry concept was invented by Bruno Canard and Simon Sarfati at the Pasteur Institute in Paris.[1][2] It was developed by Shankar Balasubramanian and David Klenerman of Cambridge University,[3] who subsequently founded Solexa, a company later acquired by Illumina. This sequencing method is based on reversible dye-terminators that enable the identification of single bases as they are introduced into DNA strands. It can also be used for whole-genome and region sequencing, transcriptome analysis, metagenomics, small RNA discovery, methylation profiling, and genome-wide protein-nucleic acid interaction analysis.[4][5]
Procedure
Tagmentation
The first step after DNA purification is tagmentation. Enzymes called transposases randomly cut the DNA into short segments ("tags"). Adapters are added on either side of the cut points (ligation). Strands that fail to have adapters ligated are washed away.[6]
Double stranded DNA is cleaved by transposomes. The cut ends are repaired and adapters, indices, primer binding sites, and terminal sites are added to each strand of the DNA. Image based in part on illumina's sequencing video[6]
Reduced cycle amplification
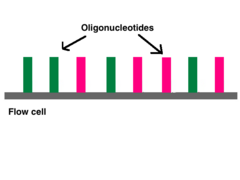
The next step is called reduced cycle amplification. During this step, sequences for primer binding, indices, and terminal sequences are added. Indices are usually six base pairs long and are used during DNA sequence analysis to identify samples. Indices allow for up to 96 different samples to be run together. During analysis, the computer will group all reads with the same index together.[7][8]The terminal sequences are used for attaching the DNA strand to the flow cell. Illumina uses a "sequence by synthesis" approach.[8]This process takes place inside of an acrylamide-coated glass flow cell.[9] The flow cell has oligonucleotides (short nucleotide sequences) coating the bottom of the cell, and they serve to hold the DNA strands in place during sequencing. The oligos match the two kinds of terminal sequences added to the DNA during reduced cycle amplification. As the DNA enters the flow cell, one of the adapters attaches to a complementary oligo.
Millions of oligos line the bottom of each flow cell lane.
Bridge amplification
Once attached, cluster generation can begin. The goal is to create hundreds of identical strands of DNA. Some will be the forward strand; the rest, the reverse. Clusters are generated through bridge amplification. Polymerases move along a strand of DNA, creating its complementary strand. The original strand is washed away, leaving only the reverse strand. At the top of the reverse strand there is an adapter sequence. The DNA strand bends and attaches to the oligo that is complementary to the top adapter sequence. Polymerases attach to the reverse strand, and its complementary strand (which is identical to the original) is made. The now double stranded DNA is denatured so that each strand can separately attach to an oligonucleotide sequence anchored to the flow cell. One will be the reverse strand; the other, the forward. This process is called bridge amplification, and it happens for thousands of clusters all over the flow cell at once.
Clonal amplification
Over and over again, DNA strands will bend and attach to oligos. Polymerases will synthesize a new strand to create a double stranded segment, and that will be denatured so that all of the DNA strands in one area are from a single source (clonal amplification). Clonal amplification is important for quality control purposes. If a strand is found to have an odd sequence, then scientists can check the reverse strand to make sure that it has the complement of the same oddity. The forward and reverse strands act as checks to guard against artifacts. Because Illumina sequencing uses polymerases, base substitution errors have been observed,[10] especially at the 3' end.[11] Paired end reads combined with cluster generation can confirm an error took place. The reverse and forward strands should be complementary to each other, all reverse reads should match each other, and all forward reads should match each other. If a read is not similar enough to its counterparts (with which it should be a clone), an error may have occurred. A minimum threshold of 97% similarity has been used in some labs' analyses.[11]
Sequence by synthesis
At the end of clonal amplification, all of the reverse strands are washed off the flow cell, leaving only forward strands. Primers attach to the forward strands and a polymerase adds fluorescently tagged nucleotides to the DNA strand. Only one base is added per round. A reversible terminator is on every nucleotide to prevent multiple additions in one round. Using the four-colour chemistry, each of the four bases has a unique emission, and after each round, the machine records which base was added. Starting with the launch of the NextSeq and later the MiniSeq, Illumina introduced a new two-colour sequencing chemistry. Nucleotides are distinguished by either one of two colours (red or green), no colour ("black") or binding both colours (appearing orange as a mixture between red and green).
Tagged nucleotides are added in order to the DNA strand. Each of the four nucleotides have an identifying label that can be excited to emit a characteristic wavelength. A computer records all of the emissions, and from this data, base calls are made.
Once the DNA strand has been read, the strand that was just added is washed away. Then, the index 1 primer attaches, polymerizes the index 1 sequence, and is washed away. The strand forms a bridge again, and the 3' end of the DNA strand attaches to an oligo on the flow cell. The index 2 primer attaches, polymerizes the sequence, and is washed away.
A polymerase sequences the complementary strand on top of the arched strand. They separate, and the 3' end of each strand is blocked. The forward strand is washed away, and the process of sequence by synthesis repeats for the reverse strand.
Data analysis
The sequencing occurs for millions of clusters at once, and each cluster has ~1,000 identical copies of a DNA insert.[10] The sequence data is analyzed by finding fragments with overlapping areas, called contigs, and lining them up. If a reference sequence is known, the contigs are then compared to it for variant identification.
This piecemeal process allows scientists to see the complete sequence even though an unfragmented sequence was never run; however, because Illumina read lengths are not very long[11] (HiSeq sequencing can produce read lengths around 90 bp long[7]), it can be a struggle to resolve short tandem repeat areas.[7][10] Also, if the sequence is de novo and so a reference doesn't exist, repeated areas can cause a lot of difficulty in sequence assembly.[10] Additional difficulties include base substitutions (especially at the 3' end of reads[11]) by inaccurate polymerases, chimeric sequences, and PCR-bias, all of which can contribute to generating an incorrect sequence.[11]
Comparison with other sequencing methods
This technique offers a number of advantages over traditional sequencing methods such as Sanger sequencing. Due to the automated nature of Illumina dye sequencing it is possible to sequence multiple strands at once and gain actual sequencing data quickly. Additionally, this method only uses DNA polymerase as opposed to multiple, expensive enzymes required by other sequencing techniques (i.e. pyrosequencing).[12]
Nanopore sequencing
From Wikipedia, the free encyclopedia
On the left is a drawing of the complex formed between alpha-hemolysin and dsDNA with linkage through an oligomer. On the right, movement of this complex in relation to a nanopore channel is shown sequentially in two steps (I) and (II). Once the complex is inserted into the nanopore, the alpha-hemolysin protein will be functional in the newly formed hybrid, biological and solid state, nanopore system.
Nanopore sequencing is a third generation[1] approach used in the sequencing of biopolymers- specifically, polynucleotides in the form of DNA or RNA.
Using nanopore sequencing, a single molecule of DNA or RNA can be sequenced without the need for PCR amplification or chemical labeling of the sample. At least one of these aforementioned steps is necessary in the procedure of any previously developed sequencing approach. Nanopore sequencing has the potential to offer relatively low-cost genotyping, high mobility for testing, and rapid processing of samples with the ability to display results in real-time. Publications on the method outline its use in rapid identification of viral pathogens,[2] monitoring ebola,[3]environmental monitoring,[4] food safety monitoring, human genome sequencing,[5] plant genome sequencing,[6] monitoring of antibiotic resistance,[7] haplotyping[8] and other applications.
Principles for detection and base identification
Nanopore sequencing uses electrophoresis to transport an unknown sample through an orifice of 10−9 meters in diameter. A nanopore system always contains an electrolytic solution- when a constant electric field is applied, an electric current can be observed in the system. The magnitude of the electric current density across a nanopore surface depends on the nanopore's dimensions and the composition of DNA or RNA that is occupying the nanopore. Sequencing is made possible because, when close enough to nanopores, samples cause characteristic changes in electric current density across nanopore surfaces. The total charge flowing through a nanopore channel is equal to the surface integral of electric current density flux across the nanopore unit normal surfaces between times t1 and t2.
Challenges
One challenge for the 'strand sequencing' method was in refining the method to improve its resolution to be able to detect single bases. In the early papers methods, a nucleotide needed to be repeated in a sequence about 100 times successively in order to produce a measurable characteristic change. This low resolution is because the DNA strand moves rapidly at the rate of 1 to 5μs per base through the nanopore. This makes recording difficult and prone to background noise, failing in obtaining single-nucleotide resolution. The problem is being tackled by either improving the recording technology or by controlling the speed of DNA strand by various protein engineering strategies and Oxford Nanopore employs a 'kmer approach', analyzing more than one base at any one time so that stretches of DNA are subject to repeat interrogation as the strand moves through the nanopore one base at a time.[28] Various techniques including algorithmic have been used to improve the performance of the MinION technology since it was first made available to users.[29] More recently effects of single bases due to secondary structure or released mononucleotides have been shown.[30][31]
Professor Hagan Bayley proposed in 2010 that creating two recognition sites within an alpha hemolysin pore may confer advantages in base recognition.[32]
One challenge for the 'exonuclease approach',[33] where a processive enzyme feeds individual bases, in the correct order, into the nanopore, is to integrate the exonuclease and the nanopore detection systems. In particular,[34] the problem is that when an exonuclease hydrolyzes the phosphodiester bonds between nucleotides in DNA, the subsequently released nucleotide is not necessarily guaranteed to directly move into, say, a nearby alpha-hemolysin nanopore. One idea is to attach the exonuclease to the nanopore, perhaps through biotinylation to the beta barrel hemolysin.[34] The central pore of the protein may be lined with charged residues arranged so that the positive and negative charges appear on opposite sides of the pore. However, this mechanism is primarily discriminatory and does not constitute a mechanism to guide nucleotides down some particular path.